Jmol
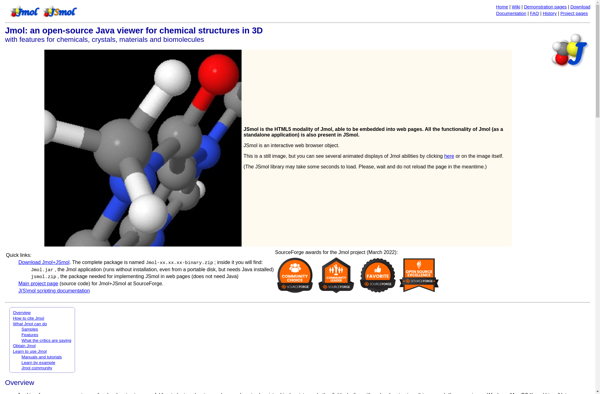
Jmol: Open-Source Java Viewer for 3D Chemical Structuress
Jmol is an open-source Java viewer for 3D chemical structures that allows users to visualize, manipulate, and analyze molecular models. It supports a variety of chemical file formats and can be used for education, research, and commercial purposes.
What is Jmol?
Jmol is an open-source molecular viewer and visualization software for 3D chemical structures, with features for chemical education, research, and commercial use. It is written in Java so that it can run on desktops and servers, online or locally. Jmol allows users to visualize, animate, manipulate, measure, and analyze a wide variety of molecular file formats including PDB, Mol2, SDF, Gaussian outputs, XYZ, and more.
Some key capabilities of Jmol include:
- Interactive visualization and smooth real-time rotation of molecular models
- Support for displaying crystal morphologies, surfaces, vibrations and orbitals
- Measuring bond lengths, angles, torsions
- Displaying space-filling, sticks, balls and sticks, ribbons, cartoon models
- Selecting, hiding and coloring atoms
- Annotating models with text labels
- Exporting high resolution image files
Jmol scripts can be embedded in web pages to create custom interactive visualizations. It has extensive documentation and tutorials available. Jmol is used worldwide for education, training, and research in fields like chemistry, biochemistry, physics, nanotechnology, and material science.
Jmol Features
Features
- 3D visualization of molecules and chemical structures
- Support for common chemical file formats like PDB, Mol, SDF
- Interactive manipulation and measurement of molecular models
- Analysis tools like molecular orbitals, electrostatics, etc
- Scripting and programming interface for automation and customization
- Platform independent (Java-based)
- Integration with web pages and applications
Pricing
- Open Source
- Free
Pros
Cons
Official Links
Reviews & Ratings
Login to ReviewThe Best Jmol Alternatives
View all Jmol alternatives with detailed comparison →
Top Science & Education and Molecular Modeling and other similar apps like Jmol
Here are some alternatives to Jmol:
Suggest an alternative ❐PyMOL
Rasmol
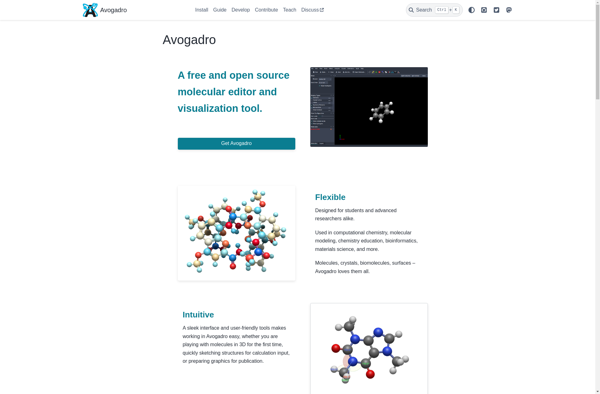
Avogadro
Ghemical

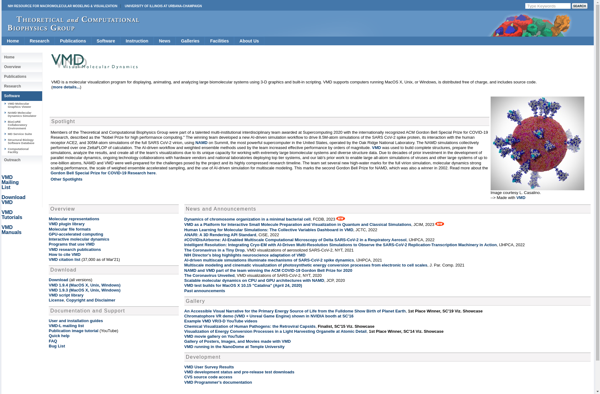
VMD - Visual Molecular Dynamics
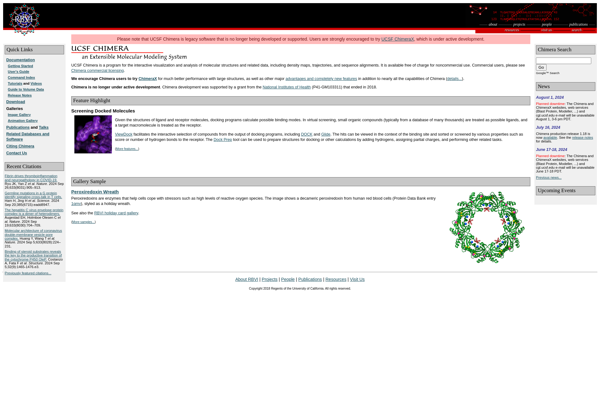
UCSF Chimera
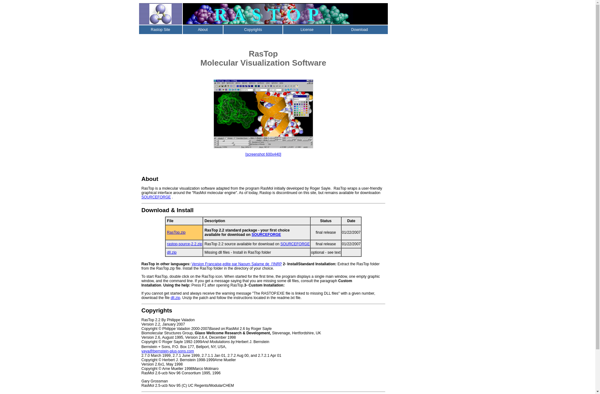
RasTop
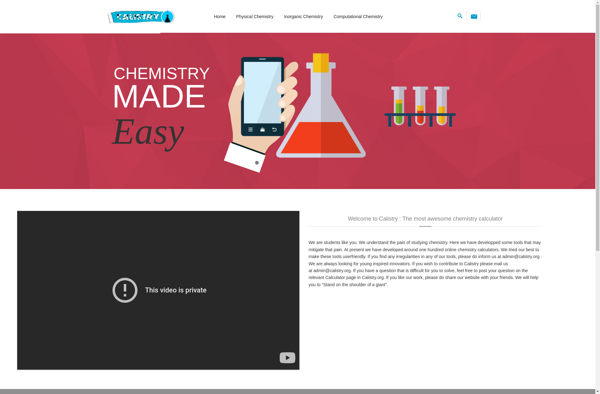
Calistry.org